Capturing carbon degrading microbes with fluorescent substrate bait
PIs: Tanja Woyke, Natalia Ivanova, Steve Singer
Biogeochemical processes in soil are greatly governed by soil organisms, including bacteria and fungi. Microbial communities are fundamental to soil C cycling, by diverse processes such as the conversion of dead plant material and organic compounds released by plant roots into long-term storage pools or CO2. Thus a knowledgebase that extends deeply into soil microbial community composition and function is urgently needed. Because the fluxes of carbon, nitrogen and other elements between plants, soil and the atmosphere depend on complex consortia of microbes, most of which are uncultivated, microbially mediated processes in soil must be studied at the community level. While both metagenomics and single-cell genomics are today’s most commonly used cultivation-independent methodologies to gain genomic insights into complex environments, they represent two extremes with complementary strengths and limitations. Single-cell genomics is laborious and will only allow the capture of a minute fraction of the existing diversity in soil, while metagenome shotgun sequencing, proper assembly and efficient binning are intractable, even at terabase scale sequencing efforts. Most importantly, neither approach is able to pair genomic data with phenotypic information about specific community members or identify and characterize organisms and pathways of particular interest, unless they are randomly encountered. This is critical, as only some taxa in soil represent the primary drivers for key biogeochemical processes, as can be exemplified by the root endophyte community. A main focus of this proposal will be the experimental capture, prior to sequencing, of specific soil microbes that catalyze a function of interest, namely plant C decomposition.
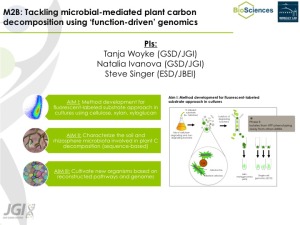
Hypothesizing that distinct soil taxa specialize in the decomposition of specific plant C substrates, we will apply novel approaches to experimentally capture bacteria that degrade plant-derived biopolymers (ie. cellulose) for sequence-based characterization. We will couple the use of fluorescently labeled substrates with single-cell sorting to obtain (i) substrate-enriched soil microbiomes for marker gene profiling and shotgun metagenomics and (ii) single-cell genomics. The approach will be developed and tested in the laboratory setting using cultivated bacterial isolates, then adopted for environmental samples and finally applied to study microbial plant C decomposition in pygmy forest soil and rhizosphere.
Reducing soil microbial diversity and complexity by selectively targeting microbes with desired functions will allow us to focus in on microbes that would otherwise be obscured and establish critically needed links between phylogeny and function. The results of this proposed work will greatly advance our understanding in soil carbon cycling and will have implications on belowground carbon storage and atmospheric C flux.